Awhile back I was piqued by a discussion on BioStar about “Where would you host your open source code repository today?“, which got me thinking about the relative merits of the different sites for hosting bioinformatics software. I am not an evangelist for any particular version control system or hosting site, and I leave it to readers to have a look into these systems themselves or at the BioStar thread for more on the relative merits of major hosting services, such as Sourceforge, Google Code, github and bitbucket. My aim here is not to advocate any particular system (although as a lab head I have certain predilections*), but to answer the straightforward empirical question: where do bioinformaticians host their code?
To do this, I’ve queried PubMed for keywords in the URLs of the four major hosting services listed above to get estimates of their uptake in biomedical publications. This simple analysis clearly has some caveats, including the fact that many publications link to hosting services in sections of the paper outside the abstract, and that many bioinformaticians (frustratingly) release code via insitutional or personal webpages. Furthermore, the various hosting services arose at different times in history, so it is also important to interpret these data in a temporal context. These (and other caveats) aside, the following provides an overview of how the bioinformatics community votes with their feet in terms of hosting their code on the major repository systems…
First of all, the bad news: of the many thousands of articles published in the field of bioinformatics, as of July Dec 31 2012 just under 700 papers (n=676) have easily discoverable code linked to a major repository in their abstract. The totals for each repository system are: 446 Sourceforge, 152 on Google Code, 78 on github and only 5 on bitbucket. So, by far, the majority of authors have chosen not to host their code on a major repository. But for the minority of authors who have chosen to release their code via a stable repository system, most use Sourceforge (which was is the oldest and most established source code repository) and effectively nobody is using bitbucket.
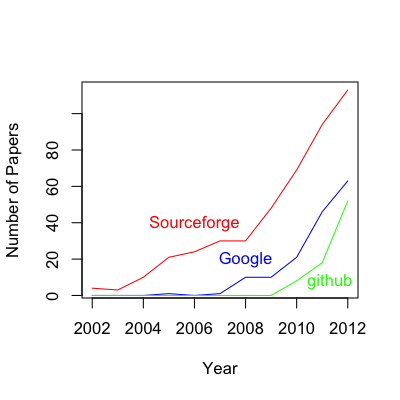
The first paper to link published code to a major repository system was only a decade ago in 2002, and a breakdown of the growth in code hosting since then looks like this:
| Year | Sourceforge | github | |
| 2002 | 4 | 0 | 0 |
| 2003 | 3 | 0 | 0 |
| 2004 | 10 | 0 | 0 |
| 2005 | 21 | 1 | 0 |
| 2006 | 24 | 0 | 0 |
| 2007 | 30 | 1 | 0 |
| 2008 | 30 | 10 | 0 |
| 2009 | 48 | 10 | 0 |
| 2010 | 69 | 21 | 8 |
| 2011 | 94 | 46 | 18 |
| 2012 | 113 | 63 | 52 |
| Total | 446 | 152 | 78 |
A few things are clear from these results: 1) there is an upward trend in biomedical researchers hosting their code on major repository sites (the apparent downturn in 2012 is because data for this year is incomplete), 2) Sourceforge has clearly been the dominant players in the biomedical code repository game to date, but 3) the current growth rate of github appears to be outstripping both Sourceforge and Google Code. Furthermore, it appears that github is not experiencing any lag in uptake, as was observed in the 2002-2004 period for Sourceforge and 2006-2009 period for Google Code. It is good to see that new players in the hosting market are being accepted at a quicker rate than they were a decade ago.
Hopefully the upward trend for bioinformaticians to release their code via a major code hosting service will continue (keep up the good work, brothers and sisters!), and this will ultimately create a snowball effect such that it is no longer acceptable to publish bioinformatics software without releasing it openly into the wild.
- As a lab manager I prefer to use Sourceforge in our published work, since Sourceforge has a very draconian policy when it come to deleting projects, which prevents accidental or willful deletion of a repository. In my opinion, Google Code and (especially) github are too permissive in terms of allowing projects to be deleted. As a lab head, I see it is my duty to ensure the long-term preservation of published code above all other considerations. I am aware that there are mechanisms to protect against deletion of repositories on github and Google Code, but I would suspect that most lab heads do not utilize them and that a substantial fraction of published academic code is one click away from deletion.
Casey – I’m also wondering to what extent projects have relocated from the repository originally reported in the article. For example, BioPerl was hosted on OBF’s version control server at the time of publication, but has since moved to Github. I think this applies to other Bio* projects, too. Perhaps this has happened in sufficient numbers to further enhance the Github uptake trajectory.
Re: project deletion policy, I fully agree with you on the principle. However, with the distributed nature of git means that you can have both the social coding features of Github and the insistence on permanence of SourceForge. Sf.net supports git, and so you could simply regularly push all updates from a github-hosted repo to a Sf.net-hosted clone of the repo. That’s fairly trivial to do.
My impression is that there has been a substantial amount of the sort of migration you indicate, but it probably wouldn’t be captured here unless authors have written an update paper with the new URL. On that note, it would be great to see a BioPerl update paper, detailing all the new bells and whistles plus a link to the new github site.
I like your suggestion for live social coding in github and creating a more permanent “version of record” in Sourceforge that is linked to in the publication.
For what it’s worth, I checked for projects tagged “bioinformatics” in each repository:
sourceforge 736
Google source 1619
Github 470
Not sure that they use the same criteria for tagging nor for counting projects, though.
Interesting, this would suggest (assuming tags are represtative) that there are many projects in Google Code that have never made it to publication, or are found in publications that link to the URL in the methods/results sections but not the abstract.
More numbers: searches in Google Scholar (including full text when possible) for “bioinformatics” and:
sourceforge: 11,700
Google code: 186
github: 524
Limiting to 2011-12:
sourceforge: 2869
Google code: 81
github: 368
So this confirms there seems to be an under-representation of Google code in published projects, that Gitub is mostly recent publications, and that Sourceforge is dominant.
I had never thought about project deletion before, that sort of ephemeral-ness (is that a word?) is scary. Is there a case for snapshot archiving of code at time of publication with DRYAD or similar? Obviously not a hosting service, but a permanent .zip file is a good complement to a deletable hosting service.
I think this is wise…perhaps as a supplemental file to the paper?
I think people are generally unaware of how poor replication, data availability, and software availability is within bioinformatics and application of bioinformatic tools… I’d be willing to bet that 90% of the code NOT hosted in a major repo is dead, and 50% of the hosted code is as well. I’m thinking of starting a Bioinformatics Software Watch Web site (much like Retraction Watch)…
There has been some work on cataloguing URL decay in PubMed (e.g. http://www.ncbi.nlm.nih.gov/pubmed/18413326), and applying this approach one could test this idea specifically: is software associated with URLs from major repositories more stable than URLs from institutes/personal webpages/etc? One for Pierre (http://plindenbaum.blogspot.co.uk/2011/04/404-not-found-database-of-non.html)?
I’m curious as to whether there are any concrete examples of projects that have code being deleted by the service provider from Google Code or Github, bioinformatics related or otherwise.
On Google Code in particular, there seem to be a huge number of well intentioned projects that provide nothing but a description – no code, no wiki pages, nothing – and yet they persist. I’ve stumbled back to these empty projects on multiple occasions spanning several years and they’ve never disappeared.
Accidental deletion is always a concern, however. Best to get a student / friend / colleague / competitor to fork your Github project so there is a second copy in someone else’s account I guess (which I have in fact done for a recent project, but not really for that reason).
I am less concerned about the hosting service deleting code (and not aware of any examples of this). I am more worried about a developer deleting a project or moving the code to another site without clear forwarding details. As a lab head, I see it as my duty to make sure that a link to code in a publication is as stable as possible over time.
You are right that forking on github is the best way to preserve a repo there, but how many people actually do this as a routine procedure? Maybe this is something journals should require or perform themselves to ensure longevity of the code…
I have seen some cases where lab heads have move out from Sourceforge and Google to Github lately.
I’ve always felt source code as supplementary data should be a requirement for any software publication, but I know not everyone agrees, and “it’s more complicated than that”(tm). The idea of journals taking responsibility for source code has certainly been discussed before in other circles – for instance I believe the idea may have been thrown around as a possible feature of http://www.openresearchcomputation.com/ (RIP).
Personally I’m not sure I’d trust most journal publishers to even manage to maintain stable links or the raw data of any zip file, let alone a functional source core repository. Broken links to Supplementary Data are not uncommon.
What gives me confidence in Github is that they have a clear business model centered around maintaining source code (rather than typesetting PDFs and selling them as bundle subscriptions), although there is a real danger with a freemium model that at some point the free offering is neglected.
As far as the stability of URLs, if you want to take responsibility the most certain way is to only publish links to a domain that you control and remember to keep registered (not an institutional domain, but YMMV depending on the institution) – eg, a lab website which can then link or forward to where the source is currently hosted. Most labs won’t do this because it’s perceived as more work to maintain, and an institutional address usually looks ‘more professional’. Given that all URLs tend to break eventually, maybe it’s really searchability that counts in the long term.
On the Preservation of Published Bioinformatics Code on Github « I wish you'd made me angry earlier
Stop Hosting Data and Code on your Lab Website | Living Biology
Archiving of bioinformatics software | m's Bioinformatics
Archiving of bioinformatics software | Bioinfo Toolbox
Archiving of bioinformatics software | Blog